Modelling the molecular world we live in, one system at a time
Imagine being able to see our world at a molecular level. Wouldn’t that be amazing? But if we could, would it be useful?

My research projects
TEM β-lactamase, inhibitors & antibiotics
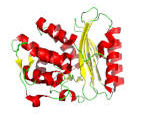
Methods development & electric fields
Green Fluorescent proteins (GFP)
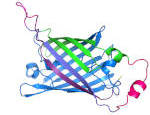
Protein engineering & enhanced sampling simulations
Liver alcohol dehydrogenase (LADH)
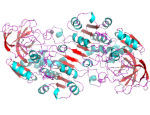
Electric fields & metal-ion enzymes
Cyclooxygenases 1 and 2 & Non-steroidal anti-inflammatory drugs (NSAID)
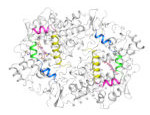
Induced-fit mechanism & selectivity, free energies & methods development
Insuline-regulated aminopeptidase (IRAP)
Drug development & binding free energies
My research
Most of traditional science is built on experiments and the results are explained using theoretical knowledge. Traditionally, drugs were discovered when people tried out new herbs or combinations of herbs and found that it made them feel better. As technology and science have progressed, traditional herbs have been purified to isolate and characterize the active substance. Sometimes, as in the case of Aspirin, the active substance has then been synthetically enhanced to improve the effect of the drug or to reduce the side-effects. However, even when a drug has been used for centuries, the mechanism of the drug can still be unknown. Knowing that a drug works is not the same as knowing how it works.
In my research, I strive to understand the dynamics of proteins and the molecular mechanisms of catalysis in and inhibition of enzymes. By combining physics-based theoretical models with computer simulations in modern high-performance computers, we can create models and make predictions. By combining simulations with experiments, new insights are obtained that can be used to develop the next generation of rationally designed drugs and molecular tools.
Do you want to join my lab?
Students
Are you a Master’s degree or ERASMUS student and want to do a degree project in my lab? Are you interested in molecular modelling, simulations, or programming and would like to contribute to antibiotic research?
Then you should contact me so we can discuss how to make it happen!
Researchers
Are you a postdoc with your own funding or a PhD student who want to do part of your project in my lab? Contact me to discuss available opportunities.
Currently I am not accepting applications for PhDs, but you might be able to do part of your project in my lab. Contact me to find out more.